Pollinators are attracted to flowers based on certain characteristics, including color, scent and morphology. Evolutionary changes in these traits correlate with changes in pollinator-plant relationships, and pollinator syndromes, or suites of floral characteristics that influence pollinator identity, can differ greatly between even closely related species. Thus, characterizing the molecular basis that underlies shifts in pollinator syndromes can lead to the discovery of speciation genes, as well as to a greater understanding of evolutionary trajectories and timelines that define the species.
A new study this week in Nature Genetics reports on a gene that controls levels of ultraviolet (UV) light absorbance in different species of Petunia, affecting whether the flowers are pollinated by bees, hawkmoths or hummingbirds. Through a series of elegant experiments involving QTL analysis, genetic crosses and a transponson mutagenesis screen, the authors were able to not only find a single gene, but also to describe the particular mutations responsible for the increased UV absorbance seen in one species and the decreased absorbance seen in another.
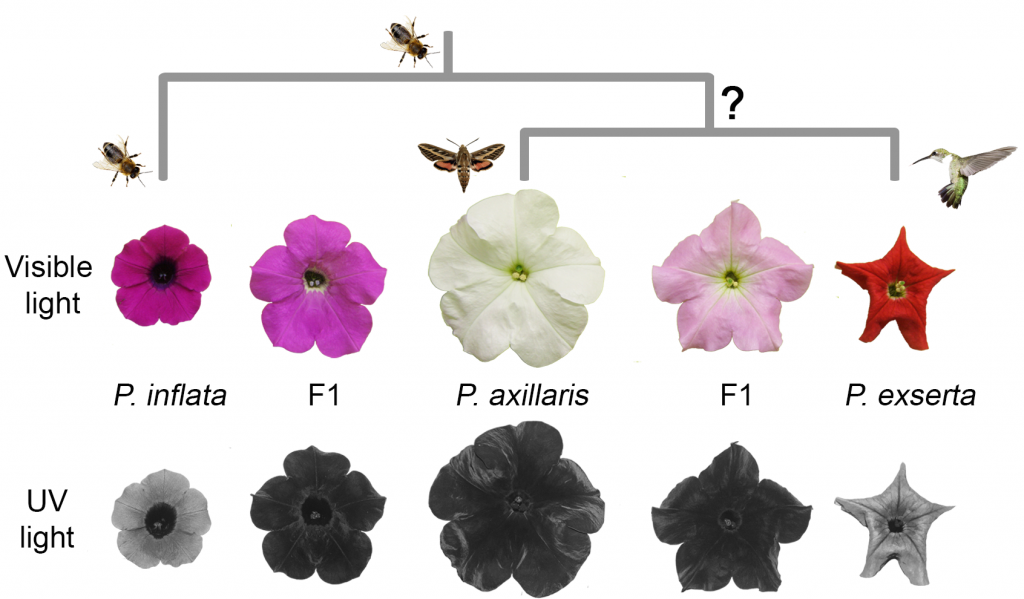
The MYB-FL gene that they isolated is a transcription factor that regulates FLS (flavonol synthase) and thus directly controls the production of flavonol, a compound that absorbs UV light. Flowers with high UV absorbance have a concomitant decrease in visible light absorbance, and this is reflected by pollinator preference. Species with low UV absorbing flowers have pink or red coloring and are pollinated by bees or hummingbirds, while species with high UV absorbing flowers have white coloring and are pollinated by (the nocturnal) hawkmoth. The authors found that the high UV absorbing species has a promoter mutation in the MYB-FL gene that increases its expression, while in the low UV absorbing species that is pollinated by hummingbirds, there is a frameshift mutation in the MYB-FL locus that compromises the function of the protein.
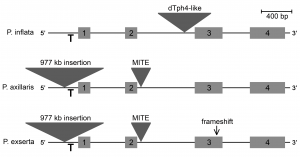
Through this analysis, the authors were able to formulate a model for the evolutionary relationships between three Petunia species. Colorful flowers that have low UV absorbance and that are bee-pollinated represent the ancestral state, as exemplified by P. inflata. The increased UV absorbance of the white flowered, hawkmoth-pollinated P. axillaris evolved via a gain-of-function cis-regulatory mutation in MYB-FL that increases its expression and thus, flavonol production. Finally, a subsequent inactivating frameshift mutation seen in P. exerta restored low UV absorbance and is associated with colorful flowers that are pollinated by hummingbirds.
We spoke with lead investigator Cris Kuhlemeier to get some background on this research.
Why do you work with Petunia? Is it a particularly good subject for studying pollination syndrome shifts?
Our goal is to find the plant genes responsible for the adaptation to different pollinators. For that, we need a system with good molecular genetics and well-defined pollination syndromes. The garden petunia has a long history as genetic model system, today it is probably best known for the discovery of RNAi. Wild Petunia species are adapted to pollination by bees, hawkmoths and hummingbirds. These species are easy to cross and propagate in the lab and give fertile offspring, and most of the genetic tools can easily be transferred from the garden petunia to the wild species.
You identified different classes of mutations in the MYB-FL gene that help to clarify evolutionary relationships between different Petunia species. What advantage does this approach have over sequencing and phylogenetic analysis?
In recent radiations such as in Petunia, classical phylogenies often have limited resolution and individual gene trees are often in conflict. We try to understand the process of adaptation and speciation by studying the gene modifications that cause reproductive isolation. By superimposing these functionally relevant polymorphisms onto the classical phylogeny, discrepancies between individual gene trees become informative.
It is interesting that you observe a trade-off between levels of anthocyanins and flavonols in these flowers. Were you expecting to see this and were you surprised that a single locus affected both levels?
Anthocyanins and flavonols share the same precursors, so finding metabolic competition was not unexpected. We started this project on the assumption that the genetics of pollination syndromes would be relative simple. At least simple enough to be able to clone the relevant genes. That a single gene can change two traits simultaneously was better than we had hoped for.
You hypothesize that R2R3-MYB transcription factors provide the toolbox for shifts in floral pollination syndromes. Do you think that your results are generalizable to other plants and/or complex traits?
R2R3-MYBs appear indeed to be over-represented, in the same way that HOX factors are overrepresented in segmentation or MADS box factors in floral organ identity. But the sample size is still small, and it is always dangerous to extrapolate, especially in ecology and evolution.
Finally, this works represents a nice combination of laboratory and field studies. Do you enjoy collecting flowers in the wild?
Well, it did rain a lot during my visit last month. But yes, it has been a new and enjoyable for me experience to go to the field with my great Brazilian colleagues. In Brazil with its great biodiversity, I also sense the excitement that, thanks to the recent progress in sequencing technology, we are no longer limited to model systems but can study interesting biological processes in almost any plant species.